Researchers Unveil a Detailed Molecular Map of Mouse Lung Development
A recent study conducted by the Guangzhou Institutes of Biomedicine and Health,Chinese Academy of Sciences (GIBH, CAS), in collaboration with the University of Science and Technology of China, Guangzhou Medical University,Guangzhou National Laboratory, and Hong Kong Institute of Science&Innovation, Chinese Academy of Sciences, was published in Science Bulletin. Researchers have created a comprehensive spatiotemporal atlas of the developing mouse lung, providing unprecedented insights into the molecular and cellular processes that shape this vital organ. The study offers a detailed map of gene expression across different stages of lung development, from embryonic day 12.5 (E12.5) to postnatal day 0 (P0).
The lung is a complex organ essential for life, responsible for gas exchange and constantly exposed to environmental hazards. Understanding its development is crucial for addressing respiratory diseases. The new study uses high-throughput spatial transcriptomics to map the gene expression patterns in developing mouse lungs. The researchers identified 10 distinct spatial domains, each corresponding to different anatomical structures and cell types within the lung.
The study reveals how the lung's airways develop along a proximal-distal axis, with distinct gene expression patterns marking the proximal (closer to the trachea) and distal (towards the alveoli) regions. Genes like Sox2 and Foxj1 are enriched in the proximal airways, while Sox9 and Etv5 are more prominent in the distal regions.
The researchers discovered two distinct alveolar niches (D2 and D7) with different maturation states. D2, marked by higher expression of genes like Angpt2 and Epha3, appears to be more mature and plays a crucial role in alveolar development around birth. They identified key transcription factors and regulatory networks driving lung development. For example, Foxa1 is essential for proximal airway development, while Tbx2 and Cux1 are critical for distal airway and alveolar maturation.They also found that signaling pathways like VEGF, ANGPT, and EPHA are highly active in the more mature alveolar niches, suggesting their role in alveolar maturation and angiogenesis.
This detailed molecular atlas provides a foundation for understanding human lung development and could lead to new therapeutic strategies for respiratory diseases. By comparing the spatial gene expression patterns in mouse and human lungs, researchers can identify conserved and species-specific features, potentially uncovering new targets for treating conditions like idiopathic pulmonary fibrosis (IPF) and chronic obstructive pulmonary disease (COPD).
While the study provides a comprehensive map of lung development up to birth, further research is needed to explore the molecular mechanisms beyond this stage. Integrating functional assays and advanced imaging techniques could offer deeper insights into the dynamic processes shaping the lung's intricate structure.
This work not only enhances our understanding of lung biology but also paves the way for innovative approaches to lung regeneration and disease treatment.
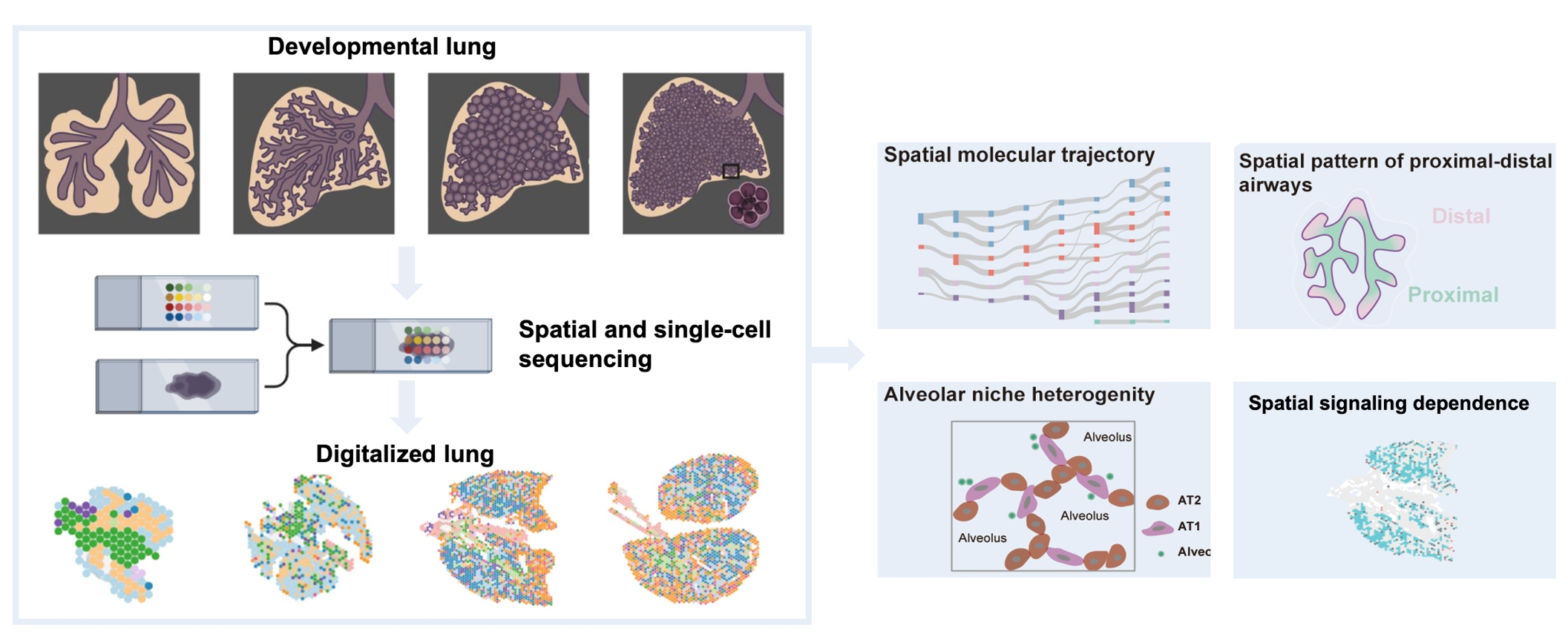
Fig. The study utilized spatial transcriptomics (10X Visium) to map gene expression from embryonic day 12.5 (E12.5) to postnatal day 0 (P0). This graphical abstract illustrates the comprehensive spatiotemporal molecular atlas of the developing mouse lung, highlighting the molecular trajectories, the spatial patterns of proximal-distal airways, alveolar niche heterogeneity, and the spatial focus of lung diseases. (Image by Prof. PENG's team)
Contacts:
PENG Guangdun, Ph.D., Principal Investigator;
Guangzhou Institutes of Biomedicine and Health, Chinese Academy of Sciences, Guangzhou, China, 510530.
Email: peng_guangdun@gibh.ac.cn
Attachment Download:
-
ContactPENG Guangdun, Ph.Dpeng_guangdun@gibh.ac.cn
-
Reference
-
Related Article